This dataset accompanies a manuscript published in Cell Reports:
Tuong, Z. K., Loudon, K. W., Berry, B., Richoz, N., Jones, J., Tan, X., Nguyen, Q., George, A., Hori, S., Field, S., Lynch, A. G., Kania, K., Coupland, P., Babbage, A., Grenfell, R., Barrett, T., Warren, A. Y., Gnanapragasam, V., Massie, C., and Clatworthy, M. R. (2021). Resolving the immune landscape of human prostate at a single-cell level in health and cancer. Cell Reports 37, 110132. 10.1016/j.celrep.2021.110132.
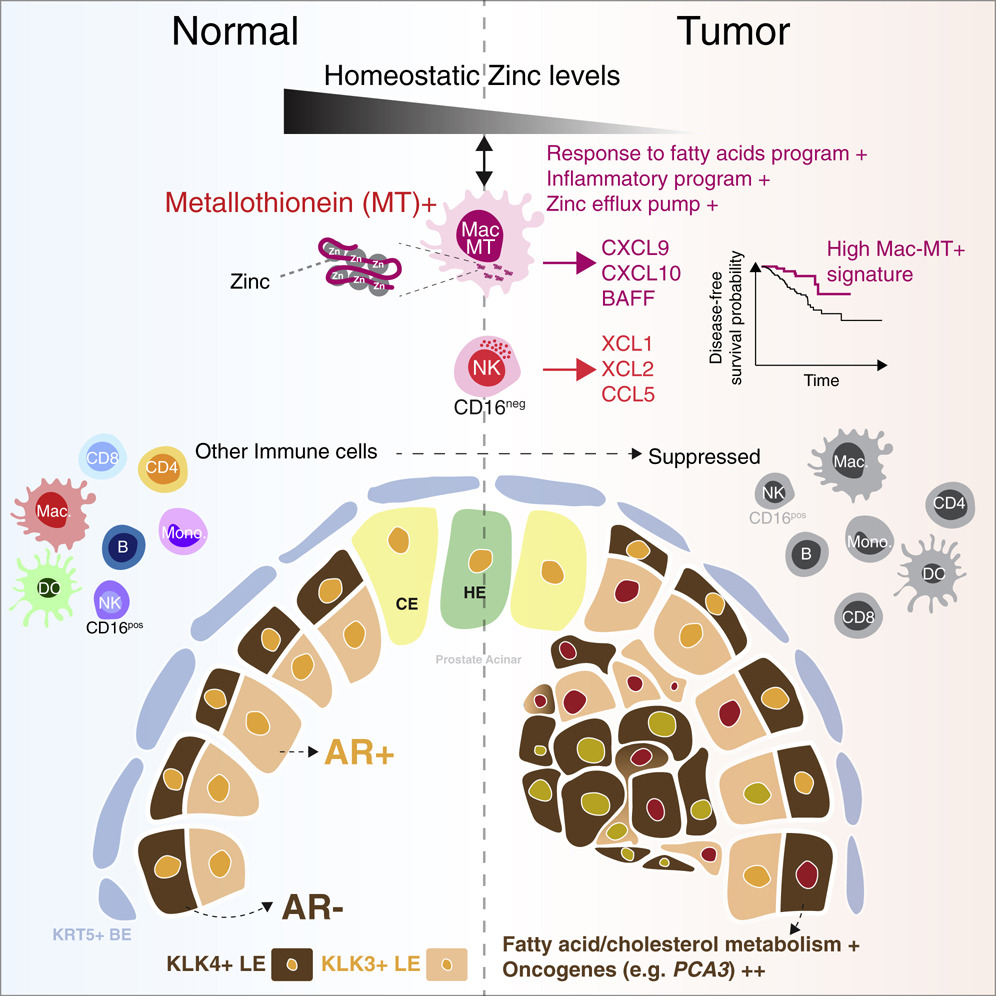
The prostate gland produces prostatic fluid, high in zinc and citrate and essential for the maintenance of spermatozoa. Prostate cancer is a common condition with limited treatment efficacy in castration-resistant metastatic disease, including with immune checkpoint inhibitors.
We used single-cell RNA-sequencing to perform an unbiased assessment of the cellular landscape of human prostate and identified a previously unappreciated subset of tumour-enriched androgen receptor-negative luminal epithelial cells, with increased expression of cancer-associated genes.
We also found a variety of innate and adaptive immune cells in normal prostate that were transcriptionally perturbed in prostate cancer. An exception was a unique, prostate-specific, zinc transporter-expressing macrophage population (MAC-MT), that contributed to tissue zinc accumulation in homeostasis, but showed enhanced inflammatory gene expression in tumours, including T cell-recruiting chemokines. Remarkably, enrichment of the MAC-MT signature in cancer biopsies was associated with improved disease-free survival, suggesting beneficial anti-tumour functions. The raw single-cell RNA-sequencing data reported in this paper is deposited at the European Genome-Phenome Archive under the accession id EGAS00001005787 with restricted data access control and will be made available by the data access committee upon reasonable request.
All code used for the study are available at
https://github.com/clatworthylab/prostateimmuneatlas
https://github.com/clatworthylab/prostateimmuneatlas
Further information and requests for resources and reagents should be directed to and will be fulfilled by the Lead Contacts, Menna R. Clatworthy (mrc38@cam.ac.uk) and Charlie Massie (cem45@cam.ac.uk)
prostate_portal_300921
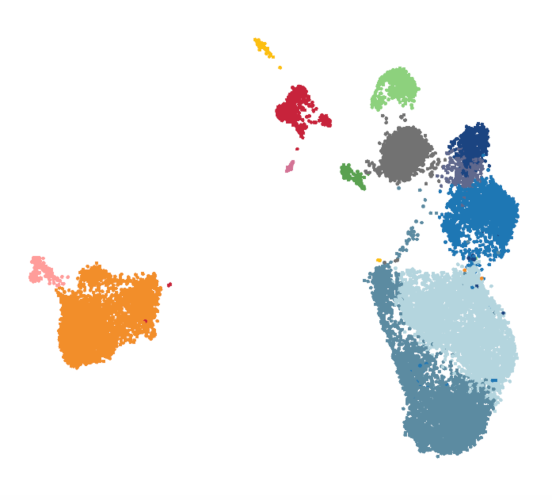

myeloid_integrated_230621